STRAP: Interactive Structure based Sequences Alignment Program.JalView documentation (Version 2) (PDF for download).JalView 1.8 at Dundee (discontinued, go to JalView 2).ESPript at IBCP Lyon: Tool to print a multiple alignment.CINEMA 5 / Utopia at Manchester (old website).CINEMA 2.1 at Manchester: Color INteractive Editor for Multiple Alignments (incl.Bork's alignment tools to enhance the results of multiple alignments (including consensus building).AMAS at Dundee: Analyse Multiply Aligned Sequences.AMAS at EBI: Analyse Multiply Aligned Sequences (OUTDATED).Software for representation of a multiple sequence alignment: Software for representation of a pairwise sequence alignment: HELP to proper use of similarity matrices.BLOSUM62: Blocks Substitution- Matrix (Henikoff and Henikoff, 1992).PAM250: Percent Accepted Mutation-Matrix (Dayhoff et al., 1978).Example of a multiple sequence alignment in Clustal and MSF format.Example of a multiple sequence alignment in MSF format.Example of a pairwise sequence alignment in MSF format.Example of a SwissProt entry in FASTA format.GRAPe at University of Oxford: Probabilistic whole-genome re-alignment (for download).FFAS03: Fold and Function Assignment System.COMPASS at University of Texas: COmparison of Multiple Protein Alignments with Assessment of Statistical Significance.COACH at Drive5: COmparison of Alignments by Constructing HMMs (software for download).SSMAL at DKFZ: Shuffled Similarities with Multiple ALignments (software for download) (OUTDATED).SATCHMO at Drive5: Simultaneous Alignment and Tree Construction using Hidden Markov m Odels (software for download).SAGA: Sequence Alignment by Genetic Algorithm (software for download).PROBCONS: Probabilistic Consistency-based Multiple Alignment of Amino Acid Sequences, at Stanford University.PRALINE: PRofile ALIg NEment, at Vrije Universiteit Amsterdam.MUSCLE at Drive5: MUltiple Sequence Comparison by Log- Expectation.MUSCA at IBM: Multiple sequence alignment using pattern discovery (only proteins!).MSA at Genestream: Multiple Sequence Alignment.Match-Box at University of Namur (only proteins!).MAFFT at EBI: Multiple Alignment using Fast Fourier Transform.Kalign at EBI: A fast and accurate multiple sequence alignment algorithm.DIALIGN at Bielefeld University: Multiple sequence alignment based on segment-to-segment comparison.COBALT at NIH: Multiple alignment incorporating pairwise constraints.ExPASy: Sequence comparisons: ClustalW, KALIGN, MAFFT, Muscle, T-Coffee, MSA, DIALGN, Match-Box, Multalin, MUSCA (see below).BCM: Multiple sequence comparisons: ClustalW, CAP, MAP, PIMA, MSA, BLOCK MAKER, MEME, Match-Box.PRSS3 at EMBnet: evaluates the significance of a protein sequence alignment.ToPlign: Login at BioSolveIT (OUTDATED).ToPlign ( Toolbox for Protein A Lignment) at Fraunhofer: Pairwise and multiple sequence comparison.SIM4 at PBIL: Program to align cDNA and genomic DNA.SIM at SIB/ExPASy: Alignment of two protein sequences with SIM, results can be viewed with LALNVIEW.LFASTA at PBIL: Local Alignment Tool for Nucleic Sequences.LALIGN at EMBnet: Finds multiple matching subsegments in two sequences (SIM-based code).BLAST 2 at NIH: Comparison of two sequence.Wise2 PromoterWise compares two DNA sequences allowing for inversions and translocations, ideal for promoters.Wise2 Dna Block Aligner aligns two sequences under the assumption that the sequences share a number of colinear blocks of conservation separated by potentially large and varied lengths of DNA in the two sequences.Wise2 (basic) compares a protein sequence to a genomic DNA sequence, allowing for introns and frameshifting errors.EMBOSS Pairwise Alignment Algorithms (global and local).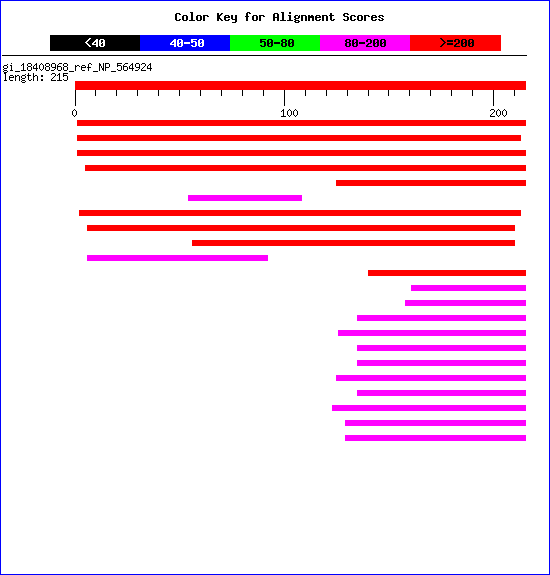 
ExPASy: Sequence comparisons: SIM, LALIGN (see below), Dotlet.BCM: Pairwise sequence comparisons: SIM, (ALIGN/LALIGN), BLAST2, LAP2, PGWISE, PCWISE.Smith & Waterman at EBI: Local alignment.Needleman & Wunsch at EBI: Global alignment.DotPlot: Dot matrix comparison of two sequences (not for Macintosh browser!).Examples of DotPlot VisualizationTechnique.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
